Expo
view channel
view channel
view channel
view channel
view channel
view channel
view channel
view channel
view channel
Clinical Chem.Molecular DiagnosticsHematologyImmunology
PathologyTechnologyIndustry
Events
Webinars

- Next Generation Automated Analyzers Increase Throughput for Clinical Chemistry and Electrolyte Testing
- Blood Metabolite Test Detects Early Cognitive Decline
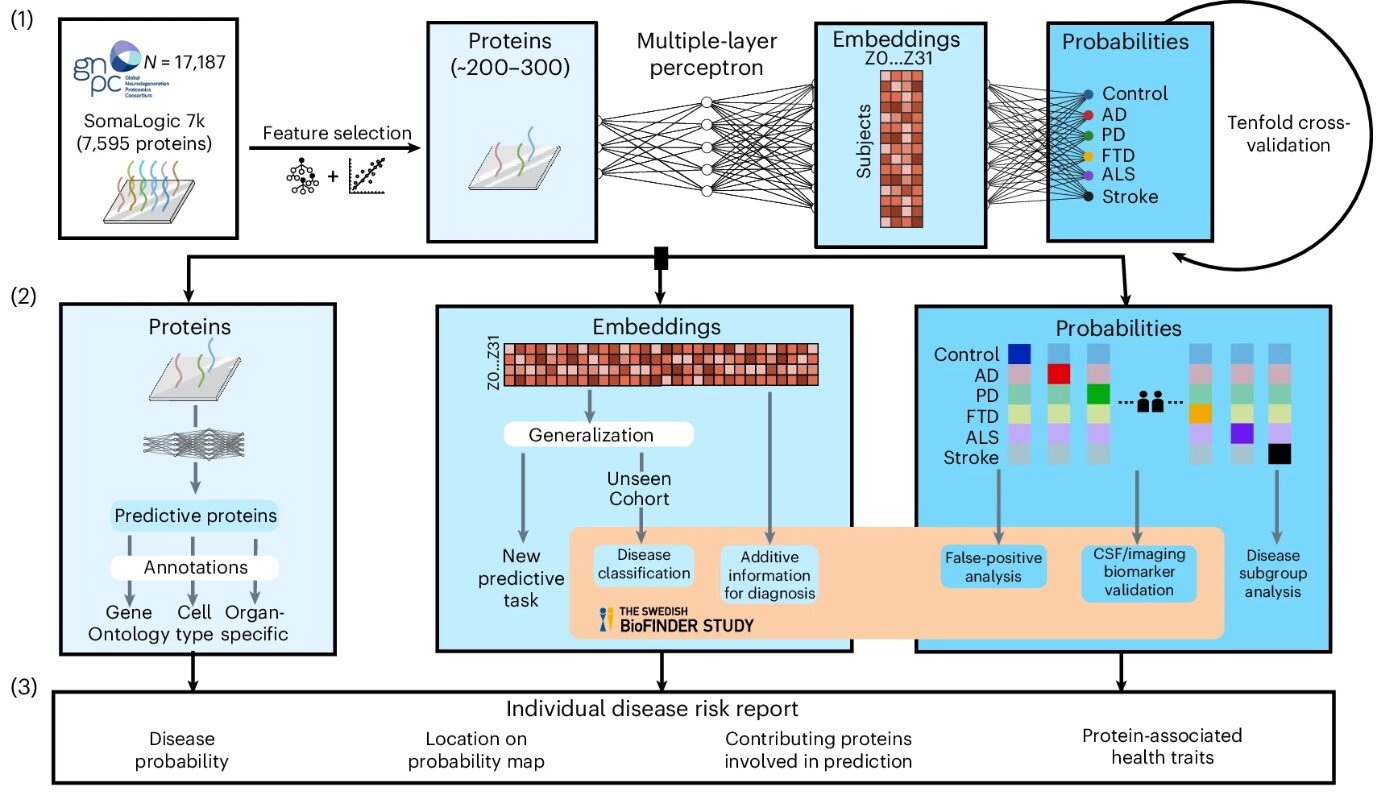
- AI-Based Blood Test Diagnose Multiple Brain Disorders from Blood Sample
- New CLIA Status Brings Mass Spectrometry Steroid Testing to Routine Labs
- Study Shows Dual Biomarkers Improve Accuracy of Alzheimer’s Detection
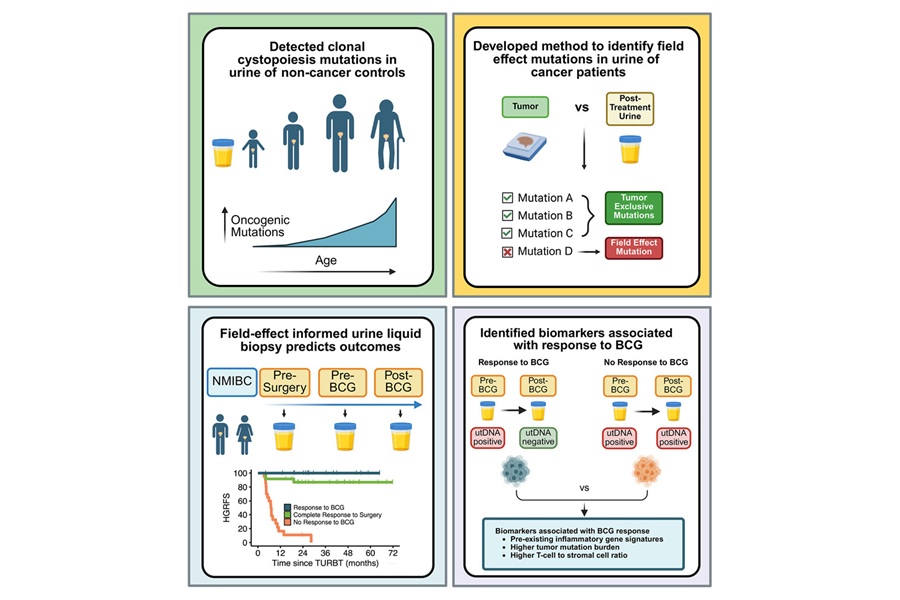
- Noninvasive Urine Test Predicts Recurrence After BCG in Bladder Cancer
- Mesothelioma in Younger Adults Linked to Genetic Risk Factors
- Immune Signatures in Blood Help Inform Cancer Risk in Lynch Syndrome
- Genetic Marker Predicts Early Heart Failure in Pulmonary Arterial Hypertension
- Genomic Subtyping Assays Identify High-Risk Early-Stage Breast Cancers
- Open Multi-Omics Platform Identifies Prognostic Subtypes in Blood Cancers
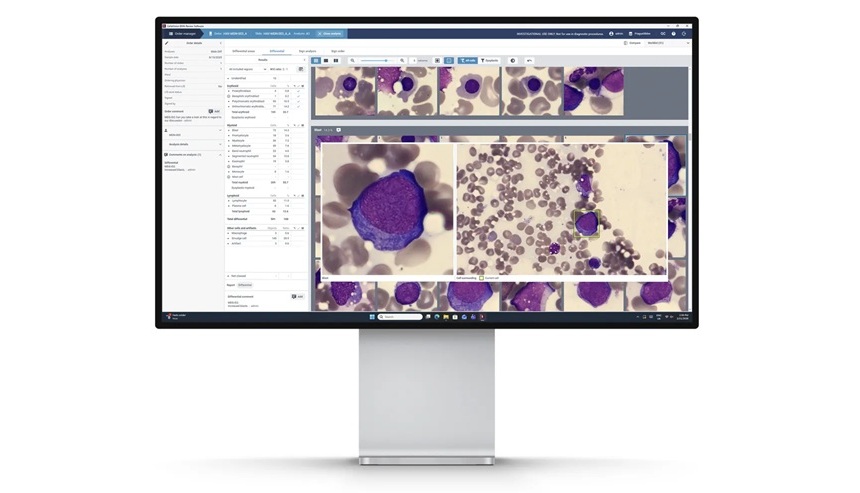
- AI-Powered Digital Workflow Standardizes Bone Marrow Aspirate Morphology
- Rapid Cartridge-Based Test Aims to Expand Access to Hemoglobin Disorder Diagnosis
- New Guidelines Aim to Improve AL Amyloidosis Diagnosis
- Fast and Easy Test Could Revolutionize Blood Transfusions
- Antibody Blood Test Identifies Active TB and Distinguishes Latent Infection
- FDA Approval Expands Use of PD-L1 Companion Diagnostic in Esophageal and GEJ Carcinomas
- Study Identifies Inflammatory Pathway Driving Immunotherapy Resistance in Bladder Cancer
- Microfluidic Chip Detects Cancer Recurrence from Immune Response Signals
- Cancer Mutation ‘Fingerprints’ to Improve Prediction of Immunotherapy Response
- New Bacterial Target Identified for Early Detection of Noma
- Genomic Analysis Links Emerging Streptococcal Strains to Specific Infections
- Rapid Urine Test Speeds Antibiotic Selection for UTIs
- WHO Endorses Rapid Point-of-Care Testing to Improve TB Detection
- Breath Analysis Approach Offers Rapid Detection of Bacterial Infection
- New AI Tool Enables Rapid Treatment Selection in Pediatric Leukemia
- Breakthrough Mass Spectrometry Design Could Enable Ultra-Low Abundance Detection
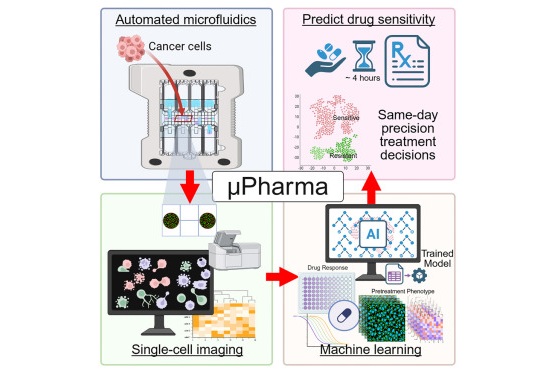
- Rapid Biosensor Detects Drug Sensitivity in Breast Tumors
- Online Tool Supports Family Screening for Inherited Cancer Risk
- Portable Breath Sensor Detects Pneumonia Biomarkers in Minutes
- Global Partnership Aims to Streamline NGS Tumor Profiling in Oncology Trials
- Takara Bio USA and Hamilton Partner Partner to Automate NGS Library Preparation
- Lunit and CellCarta Collaborate to Expand AI Pathology in CDx Development
- Integrated DNA Technologies Expands into Clinical Diagnostics
- Co-Diagnostics Agreement Expands Commercial and Distribution Reach in South Asia
- Study Links Midlife Vitamin D to Lower Tau in Alzheimer's
- Common Metablolic Enzyme Could Predict Response to Cancer Immunotherapy
- Newly Identfied Genetic Variants in MND Support Prognosis and Family Testing
- International Consensus Standardizes Tumor Microbiota Detection and Reporting
- Innate Immunity Variants Associated With Earlier Breast Cancer in BRCA1 Carriers
- Tumor-Specific Biomarker Predicts Neoadjuvant Immunotherapy Response in Gastric Cancer
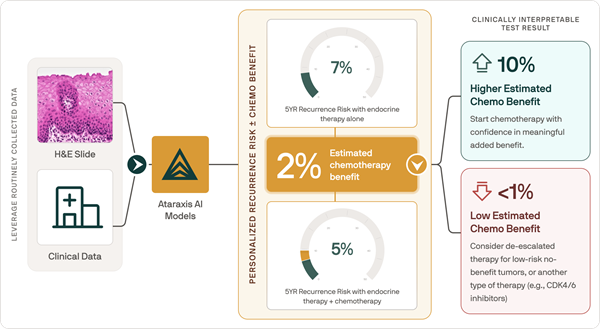
- AI Tool Predicts Patient-Specific Chemotherapy Benefit in Breast Cancer
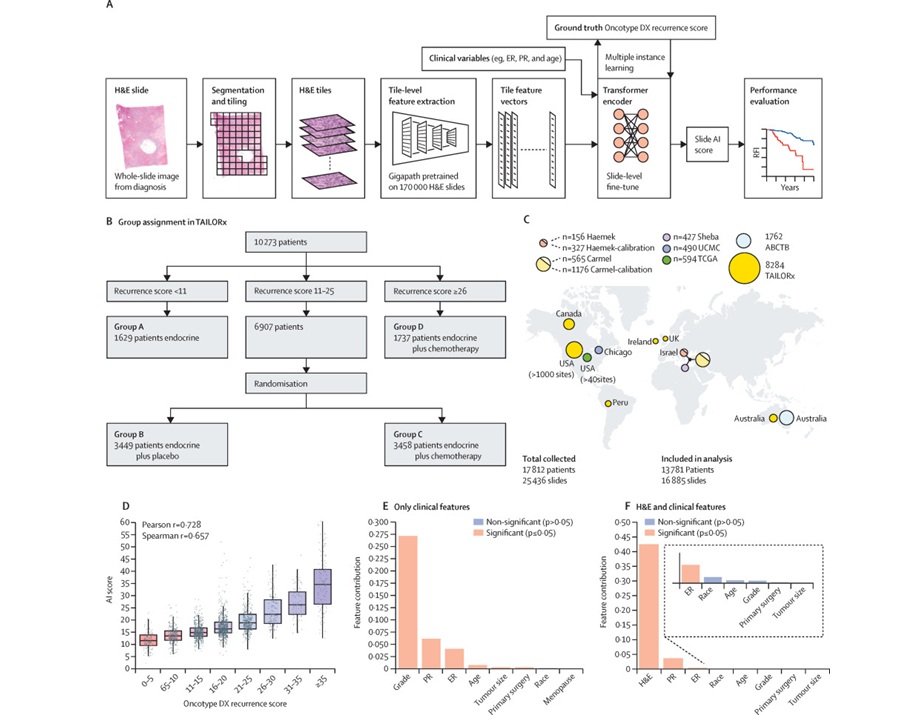
- AI-Based Pathology Model Guides Chemotherapy Decisions in Breast Cancer
- Biopsy-Based Gene Test Predicts Recurrence Risk in Lung Adenocarcinoma
- AI-Powered Tool to Transform Dermatopathology Workflow

 Expo
Expo
- Next Generation Automated Analyzers Increase Throughput for Clinical Chemistry and Electrolyte Testing
- Blood Metabolite Test Detects Early Cognitive Decline
- AI-Based Blood Test Diagnose Multiple Brain Disorders from Blood Sample
- New CLIA Status Brings Mass Spectrometry Steroid Testing to Routine Labs
- Study Shows Dual Biomarkers Improve Accuracy of Alzheimer’s Detection
- Noninvasive Urine Test Predicts Recurrence After BCG in Bladder Cancer
- Mesothelioma in Younger Adults Linked to Genetic Risk Factors
- Immune Signatures in Blood Help Inform Cancer Risk in Lynch Syndrome
- Genetic Marker Predicts Early Heart Failure in Pulmonary Arterial Hypertension
- Genomic Subtyping Assays Identify High-Risk Early-Stage Breast Cancers
- Open Multi-Omics Platform Identifies Prognostic Subtypes in Blood Cancers
- AI-Powered Digital Workflow Standardizes Bone Marrow Aspirate Morphology
- Rapid Cartridge-Based Test Aims to Expand Access to Hemoglobin Disorder Diagnosis
- New Guidelines Aim to Improve AL Amyloidosis Diagnosis
- Fast and Easy Test Could Revolutionize Blood Transfusions
- Antibody Blood Test Identifies Active TB and Distinguishes Latent Infection
- FDA Approval Expands Use of PD-L1 Companion Diagnostic in Esophageal and GEJ Carcinomas
- Study Identifies Inflammatory Pathway Driving Immunotherapy Resistance in Bladder Cancer
- Microfluidic Chip Detects Cancer Recurrence from Immune Response Signals
- Cancer Mutation ‘Fingerprints’ to Improve Prediction of Immunotherapy Response
- New Bacterial Target Identified for Early Detection of Noma
- Genomic Analysis Links Emerging Streptococcal Strains to Specific Infections
- Rapid Urine Test Speeds Antibiotic Selection for UTIs
- WHO Endorses Rapid Point-of-Care Testing to Improve TB Detection
- Breath Analysis Approach Offers Rapid Detection of Bacterial Infection
- New AI Tool Enables Rapid Treatment Selection in Pediatric Leukemia
- Breakthrough Mass Spectrometry Design Could Enable Ultra-Low Abundance Detection
- Rapid Biosensor Detects Drug Sensitivity in Breast Tumors
- Online Tool Supports Family Screening for Inherited Cancer Risk
- Portable Breath Sensor Detects Pneumonia Biomarkers in Minutes
- Global Partnership Aims to Streamline NGS Tumor Profiling in Oncology Trials
- Takara Bio USA and Hamilton Partner Partner to Automate NGS Library Preparation
- Lunit and CellCarta Collaborate to Expand AI Pathology in CDx Development
- Integrated DNA Technologies Expands into Clinical Diagnostics
- Co-Diagnostics Agreement Expands Commercial and Distribution Reach in South Asia
- Study Links Midlife Vitamin D to Lower Tau in Alzheimer's
- Common Metablolic Enzyme Could Predict Response to Cancer Immunotherapy
- Newly Identfied Genetic Variants in MND Support Prognosis and Family Testing
- International Consensus Standardizes Tumor Microbiota Detection and Reporting
- Innate Immunity Variants Associated With Earlier Breast Cancer in BRCA1 Carriers
- Tumor-Specific Biomarker Predicts Neoadjuvant Immunotherapy Response in Gastric Cancer
- AI Tool Predicts Patient-Specific Chemotherapy Benefit in Breast Cancer
- AI-Based Pathology Model Guides Chemotherapy Decisions in Breast Cancer
- Biopsy-Based Gene Test Predicts Recurrence Risk in Lung Adenocarcinoma
- AI-Powered Tool to Transform Dermatopathology Workflow


















_image.png)